import ee
import geemap.core as geemap
import pandas as pd
import pyreadr12 Subcriterion A2b
Reproducible workflow using Python
This document shows an example calculation of criterion B using data assets and functions available on Earth Engine (EE) from a notebook running Python.
12.1 Updating the assessment workflow
The original workflow included a combination of two remote sensing products (Friedl et al. 2010; Hansen et al. 2013). The updated workflow is based on an updated version of one data source that spans more years and uses improved algorithms. We have two version of the workflow, one based on R and local copies of the remote sensing products (see Section 11.1), and one based on Python and Earth Engine for cloud computing (this document).
Differences between original and updated assessment workflows for subcriterion A2b.
| Step | 2019 | 2025 (R) | 2025 (Python) |
|---|---|---|---|
| GIS data prep. | GRASS GIS / gdal / R |
R / gdal / terra |
Earth Engine |
| Data source | GFC v1.2 (2015) and MODIS LC v5.1 | GFC v1.12 (2024) | GFC v1.12 (2024) |
| Analysis | R / glm |
R / glm / redlistr |
Python / Earth Engine |
| Timeframe | 2000-2050 | 2000-2050 | 2000-2050 |
Part of the code is based on examples from Earth Engine Guides.
12.2 Set up
Import modules
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as pltAuthenticate to Earth Engine
myproject='ee-jrferrerparis'ee.Authenticate()
ee.Initialize(project=myproject)Read inputs
Here we will use three assets, two of them are available in Earth Engine’s data catalog:
gfc=ee.Image('UMD/hansen/global_forest_change_2024_v1_12')
countries=ee.FeatureCollection('USDOS/LSIB_SIMPLE/2017')The third asset has been uploaded from (FerrerParis20025?) into my personal project:
macrogroups=ee.Image('projects/{}/assets/IVC_NS_v7_270m_cea'.format(myproject))12.3 Prepare the data
First we need to select the target macrogroup, in this case the raster value is the ID, so value 563 returns the potential distribution of M563:
MG563 = macrogroups.eq(563);Then we define a region of interest, in this case we know this macrogroup is present in three countries: Colombia, Venezuela, and Trinidad and Tobago, so we will create regions for each one of them.
rVEN = countries.filter(
ee.Filter.inList('country_na', ['Venezuela',])
)
rCOL = countries.filter(
ee.Filter.inList('country_na', ['Colombia',])
)
rTT = countries.filter(
ee.Filter.inList('country_na', ['Trinidad & Tobago',])
)Parameters and functions
We set up the scale for the outputs
params = {'scale': 30, 'maxPixels':1e10}And calculate the corresponding resolution
res_km2 = (params['scale']**2)/1e6Now we design our own combined reducer for the spatial analysis:
combinedReducer = ee.Reducer.sum().combine(
reducer2 = ee.Reducer.count(),
sharedInputs = True
).combine(
reducer2 = ee.Reducer.mean(),
sharedInputs = True
)12.4 Analysis approach: Compute totals
The first approach is to calculate raster totals and do the calculations afterwards. We choose treecover, gain and loss bands from GFC and apply the mask for the potential distribution of our target macrogroup.
mcdgTotals = (
gfc
.select(['treecover2000','gain','loss'])
.selfMask()
.mask(MG563)
)Computations
We compute summaries of the three bands for each country
totalsVEN = mcdgTotals.reduceRegion(
reducer=combinedReducer,
geometry=rVEN.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()totalsVEN{'gain_count': 105206342,
'gain_mean': 0.0040003273355281,
'gain_sum': 420784.8392156863,
'loss_count': 105206342,
'loss_mean': 0.0586245309569536,
'loss_sum': 6166573.823529413,
'treecover2000_count': 105206342,
'treecover2000_mean': 37.97777763966412,
'treecover2000_sum': 3994791355.184314}totalsCOL = mcdgTotals.reduceRegion(
reducer=combinedReducer,
geometry=rCOL.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()totalsTT = mcdgTotals.reduceRegion(
reducer=combinedReducer,
geometry=rTT.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()Aggregate results
totals = pd.DataFrame([totalsVEN,totalsCOL,totalsTT])totals| gain_count | gain_mean | gain_sum | loss_count | loss_mean | loss_sum | treecover2000_count | treecover2000_mean | treecover2000_sum | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 105206342 | 0.004000 | 420784.839216 | 105206342 | 0.058625 | 6.166574e+06 | 105206342 | 37.977778 | 3.994791e+09 |
| 1 | 54502069 | 0.014495 | 789892.454902 | 54502069 | 0.103135 | 5.620141e+06 | 54502069 | 35.044587 | 1.909694e+09 |
| 2 | 233854 | 0.003176 | 741.000000 | 233854 | 0.045033 | 1.050737e+04 | 233854 | 75.694211 | 1.766134e+07 |
totals['eco_extent_2000'] = (totals.treecover2000_sum / 100) * res_km2 totals['eco_extent_2024'] = totals['eco_extent_2000'] - ((totals.loss_sum.sum()) * res_km2) + ((totals.gain_sum.sum()) * res_km2)totals[['eco_extent_2000','eco_extent_2024']]| eco_extent_2000 | eco_extent_2024 | |
|---|---|---|
| 0 | 35953.122197 | 26425.898653 |
| 1 | 17187.244420 | 7660.020876 |
| 2 | 158.952065 | -9368.271479 |
12.5 Analysis approach: Raster algebra
The second approach is to use raster algebra to calculate a new image of tree cover for 2024 that reflects pixel by pixel changes.
mcdg_2024 = (
gfc
.select(['treecover2000'])
.multiply(gfc.select(['loss']).eq(0))
.add(gfc.select(['gain']).multiply(100))
.mask(MG563)
)Computations
finalVEN = mcdg_2024.reduceRegion(
reducer=combinedReducer,
geometry=rVEN.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()finalCOL = mcdg_2024.reduceRegion(
reducer=combinedReducer,
geometry=rCOL.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()finalTT = mcdg_2024.reduceRegion(
reducer=combinedReducer,
geometry=rTT.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()Aggregate results
finals = pd.DataFrame([finalVEN,finalCOL,finalTT])finals| treecover2000_count | treecover2000_mean | treecover2000_sum | |
|---|---|---|---|
| 0 | 105206342 | 34.141011 | 3.591211e+09 |
| 1 | 54502069 | 28.821156 | 1.570559e+09 |
| 2 | 233854 | 71.958293 | 1.678966e+07 |
finals['treecover2000_sum'] / totals['treecover2000_sum'] 0 0.898973
1 0.822414
2 0.950645
Name: treecover2000_sum, dtype: float64sum(finals['treecover2000_sum']) / sum(totals['treecover2000_sum'] )0.8744396077079528253656.4 / sum(totals['gain_count'])0.001585924771041600412.6 Analysis approach: Cross tabulation
A third approach is to do a crosstabulation of tree cover loss per year, we use the bands treecover2000 and lossyear from the GFC dataset.
mcdgTC = (
gfc
.select(['treecover2000','lossyear'])
.mask(MG563)
)Computations
tclVEN = mcdgTC.reduceRegion(
reducer=combinedReducer.group(groupField=1, groupName='lossyear'),
geometry=rVEN.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()tclCOL = mcdgTC.reduceRegion(
reducer=combinedReducer.group(groupField=1, groupName='lossyear'),
geometry=rCOL.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()tclTT = mcdgTC.reduceRegion(
reducer=combinedReducer.group(groupField=1, groupName='lossyear'),
geometry=rTT.geometry(),
scale=params['scale'],
maxPixels=params['maxPixels'],
).getInfo()Crosstabulation of loss per year
Create a function to prepare the results of the previous computation for the extrapolation step:
def prepareDF(r1,r2):
# We read the data from the treecover / lossyear crosstabulation:
df = pd.DataFrame(r1['groups'])
# We use the values calculated from the gain layer to correct the area estimates:
df['gain'] = (r2['gain_sum'] / 24)
df.loc[df.lossyear==0,'gain'] = 0
# We calculate the area for each year by dividing the treecover by 100,
# adding a portion of the gain and using the resolution to transform to area:
df['tc_area'] = df['area'] = ((df['sum']/100) + df['gain']) * res_km2
# We reorder the data to account for the area lost between 2001 and 2024 and the area remaining in 2025
df['order'] = 25 - df['lossyear']
df['year'] = 1999 + df['lossyear']
df.loc[df.lossyear==0,'order'] = 0
df.loc[df.lossyear==0,'year'] = 2025
df.sort_values('order', inplace=True)
# Now calculate the accumulated area in reverse order of years:
df['tc_area'] = np.cumsum( df['area'])
# And reorder to get a time series of remaining area per year:
df.sort_values('year', inplace=True)
return(df)Apply this function to the data from Venezuela
resultsVEN = prepareDF(tclVEN,totalsVEN)resultsVEN.head()| count | lossyear | mean | sum | gain | tc_area | area | order | year | |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 352765 | 1 | 73.815910 | 2.603826e+07 | 17532.701634 | 36331.828552 | 250.123728 | 24 | 2000 |
| 2 | 224563 | 2 | 74.279556 | 1.667921e+07 | 17532.701634 | 36081.704824 | 165.892364 | 23 | 2001 |
| 3 | 217493 | 3 | 80.186956 | 1.743915e+07 | 17532.701634 | 35915.812460 | 172.731799 | 22 | 2002 |
| 4 | 188625 | 4 | 74.362437 | 1.402577e+07 | 17532.701634 | 35743.080661 | 142.011373 | 21 | 2003 |
| 5 | 241077 | 5 | 75.143395 | 1.811477e+07 | 17532.701634 | 35601.069287 | 178.812379 | 20 | 2004 |
Apply this function to the data from Colombia
resultsCOL = prepareDF(tclCOL,totalsCOL)Apply this function to the data from Trinidad and Tobago
resultsTT = prepareDF(tclTT,totalsTT)12.7 Extrapolation of decline
Xnew=pd.DataFrame(range(2000,2050),columns=['year',]).assign(intercept=1)
Xnew['year_squared'] = Xnew['year']**2def fitExtModel(df):
X = (df
.assign(intercept=1)
.reset_index(drop=True)
)
X['year_squared'] = X['year']**2
y = X.pop('tc_area')
model_1 = sm.GLM(
y,
X[['intercept', 'year', 'year_squared']],
family=sm.families.Poisson(),
)
return model_1.fit()extVEN = fitExtModel(resultsVEN)
print(extVEN.summary()) Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: tc_area No. Observations: 25
Model: GLM Df Residuals: 22
Model Family: Poisson Df Model: 2
Link Function: Log Scale: 1.0000
Method: IRLS Log-Likelihood: -159.71
Date: Mon, 25 Aug 2025 Deviance: 12.561
Time: 20:38:29 Pearson chi2: 12.6
No. Iterations: 3 Pseudo R-squ. (CS): 1.000
Covariance Type: nonrobust
================================================================================
coef std err z P>|z| [0.025 0.975]
--------------------------------------------------------------------------------
intercept -98.7564 90.558 -1.091 0.275 -276.247 78.735
year 0.1140 0.090 1.266 0.205 -0.062 0.290
year_squared -2.968e-05 2.24e-05 -1.327 0.185 -7.35e-05 1.42e-05
================================================================================prdVEN=extVEN.get_prediction(Xnew[['intercept', 'year', 'year_squared']]).summary_frame()plt.plot(resultsVEN[['year']], resultsVEN[['tc_area']], 'o')
plt.plot(Xnew[['year']], prdVEN["mean"], '--', label='fit')
plt.ylabel("Area estimated from tree cover %")
plt.xlabel("Year")
plt.fill_between(
Xnew['year'], prdVEN["mean_ci_lower"], prdVEN["mean_ci_upper"], color="k", alpha=0.1,
label='plausible range'
);
plt.legend()
plt.show()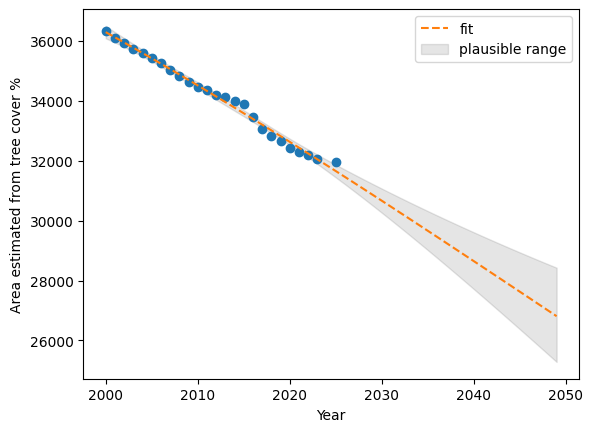
100 - (prdVEN.iloc[49][["mean","mean_ci_lower","mean_ci_upper"]].values * 100 / resultsVEN.iloc[0][['tc_area']].values)array([26.20915397, 30.41213941, 21.75231554])extCOL = fitExtModel(resultsCOL)
prdCOL=extCOL.get_prediction(Xnew[['intercept', 'year', 'year_squared']]).summary_frame()
100 - (prdCOL.iloc[49][["mean","mean_ci_lower","mean_ci_upper"]].values * 100 / resultsCOL.iloc[0][['tc_area']].values)array([52.71523956, 56.68077749, 48.38668748])plt.plot(resultsCOL[['year']], resultsCOL[['tc_area']], 'o')
plt.plot(Xnew[['year']], prdCOL["mean"], '--', label='fit')
plt.ylabel("Area estimated from tree cover %")
plt.xlabel("Year")
plt.fill_between(
Xnew['year'], prdCOL["mean_ci_lower"], prdCOL["mean_ci_upper"], color="k", alpha=0.1,
label='plausible range'
);
plt.legend()
plt.show()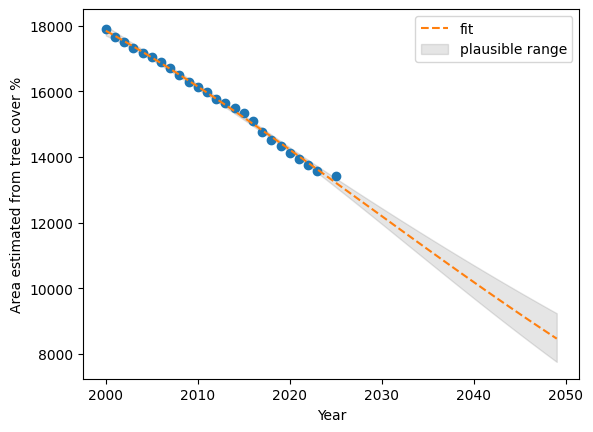
extTT = fitExtModel(resultsTT)
prdTT=extTT.get_prediction(Xnew[['intercept', 'year', 'year_squared']]).summary_frame()
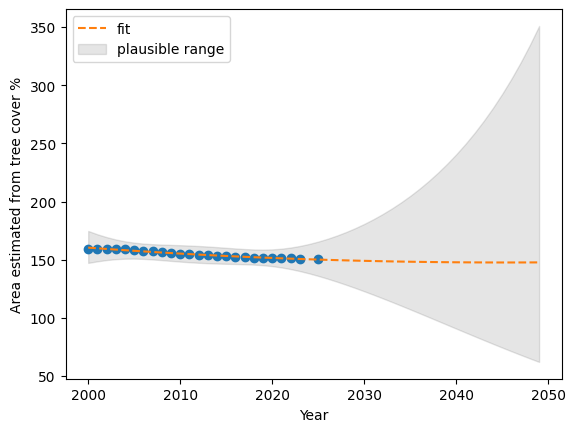
prdTT.iloc[49][["mean","mean_ci_lower","mean_ci_upper"]].values * 100 / resultsTT.iloc[0][['tc_area']].valuesarray([ 92.5565775 , 38.95877469, 219.89192697])resultsTT.iloc[0][['tc_area']].valuesarray([159.61896455])plt.plot(resultsTT[['year']], resultsTT[['tc_area']], 'o')
plt.plot(Xnew[['year']], prdTT["mean"], '--', label='fit')
plt.ylabel("Area estimated from tree cover %")
plt.xlabel("Year")
plt.fill_between(
Xnew['year'], prdTT["mean_ci_lower"], prdTT["mean_ci_upper"], color="k", alpha=0.1,
label='plausible range'
);
plt.legend()
plt.show()