import pandas as pd
import folium
from datetime import datetime, timedelta
from pyinaturalist import (
Observation,
get_observations,
get_projects_by_id,
pprint,
)
import ipyplot
from itertools import compressMy contributions to the Great Southern Bioblitz in Cairns
Cairns
Python
pyinaturalist
Folium
A summary of my iNaturalist observations during our visit to Cairns.
Load Python modules
import seaborn as sns
import matplotlib.pyplot as pltDownload iNaturalist observations
From the end of May to the first week of June 2025
projects = get_projects_by_id([253831, 255314])
pprint(projects)ID Title Type URL ━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━ 255314 Great Southern BioBlitz 2025: Cairns collection https://www.inaturalist.org/projects/255314 253831 Great Southern Bioblitz 2025: Great Barrier collection https://www.inaturalist.org/projects/253831 Reef
gbs_cairns = projects['results'][0]observations = get_observations(project_id=gbs_cairns['id'],
per_page=0)
n_obs = observations['total_results']
print("Project _{}_ has {} observations in iNaturalist".format(gbs_cairns['title'],n_obs))Project _Great Southern BioBlitz 2025: Cairns_ has 571 observations in iNaturalistrecords=list()
j=1
while len(records) < n_obs:
print("Requesting observations from project _{}_: page {}, total of {} observations downloaded".format(gbs_cairns['title'],j,min(j*200,n_obs)))
observations = get_observations(project_id=gbs_cairns['id'],per_page=200,page=j)
for obs in observations['results']:
record = {
'uuid': obs['uuid'],
'user_id': obs['user']['id'],
'user': obs['user']['login'],
'user_name': obs['user']['name'],
'user_icon': obs['user']['icon_url'],
'day_observed': obs['observed_on_details']['day'],
'species_guess': obs['species_guess'],
'quality_grade': obs['quality_grade'],
'description': obs['description'],
'location': obs['location']
}
if obs['taxon'] is not None:
if 'iconic_taxon_name' in obs['taxon'].keys():
record['iconic_taxon'] = obs['taxon']['iconic_taxon_name']
if len(obs['observation_photos'])>0:
record['url'] = obs['observation_photos'][0]['photo']['url']
record['attribution'] = obs['observation_photos'][0]['photo']['attribution']
if obs['oauth_application_id'] is None:
record['app'] = 0
else:
record['app'] = obs['oauth_application_id']
records.append(record)
j=j+1Requesting observations from project _Great Southern BioBlitz 2025: Cairns_: page 1, total of 200 observations downloaded
Requesting observations from project _Great Southern BioBlitz 2025: Cairns_: page 2, total of 400 observations downloaded
Requesting observations from project _Great Southern BioBlitz 2025: Cairns_: page 3, total of 571 observations downloadedproject_obs = pd.DataFrame(records)Map of observations
map = folium.Map(tiles="Esri NatGeoWorldMap")fg = folium.FeatureGroup(name="iNaturalist observations", control=True, attribution="observers @ iNaturalist").add_to(map)
popup_text = """<img src='{url}'>
<caption><i>{species}</i> observed on day {observed_on} / {attribution}</caption> {desc}
"""
popup_text_no_foto = """
<caption><i>{species}</i> observed on day {observed_on} / No foto</caption> {desc}
"""
for obs in records:
if obs['quality_grade'] == 'research':
if obs['description'] is None:
desc = ""
else:
desc = obs['description']
pincolor = 'green'
else:
desc = "Observation is not research quality grade."
pincolor = 'gray'
if 'url' not in obs.keys():
fg.add_child(
folium.Marker(
location=obs['location'],
popup=popup_text_no_foto.format(
species=obs['species_guess'],
observed_on=obs['day_observed'],
desc=desc),
icon=folium.Icon(color=pincolor),
)
)
else:
fg.add_child(
folium.Marker(
location=obs['location'],
popup=popup_text.format(
species=obs['species_guess'],
observed_on=obs['day_observed'],
desc=desc,
url = obs['url'],
attribution = obs['attribution']),
icon=folium.Icon(color=pincolor),
)
)
folium.LayerControl().add_to(map)<folium.map.LayerControl object at 0x12cb26de0>
map.fit_bounds(map.get_bounds())
mapMake this Notebook Trusted to load map: File -> Trust Notebook
Observations per iconic taxon
project_obs.groupby(['iconic_taxon']).agg({
'uuid': 'count',
'species_guess': ['count',pd.Series.nunique],
'user_id': pd.Series.nunique})| uuid | species_guess | user_id | ||
|---|---|---|---|---|
| count | count | nunique | nunique | |
| iconic_taxon | ||||
| Actinopterygii | 60 | 51 | 38 | 10 |
| Amphibia | 6 | 6 | 4 | 5 |
| Animalia | 28 | 26 | 18 | 10 |
| Arachnida | 47 | 46 | 36 | 14 |
| Aves | 190 | 185 | 90 | 31 |
| Fungi | 7 | 6 | 6 | 6 |
| Insecta | 104 | 88 | 77 | 28 |
| Mammalia | 2 | 2 | 2 | 2 |
| Mollusca | 16 | 13 | 12 | 5 |
| Plantae | 90 | 82 | 67 | 20 |
| Protozoa | 1 | 0 | 0 | 1 |
| Reptilia | 20 | 18 | 12 | 10 |
data_crosstab = pd.crosstab(project_obs['iconic_taxon'],
project_obs['day_observed'],
margins = False)plt.figure(figsize=(8, 6))
sns.heatmap(data_crosstab, annot=True, fmt='d', cmap='YlGnBu', linewidths=.5)
plt.title('Heatmap of observations per day of Bioblitzing')
plt.ylabel('Iconic taxon')
plt.xlabel('Day of observation')
plt.tight_layout()
plt.show()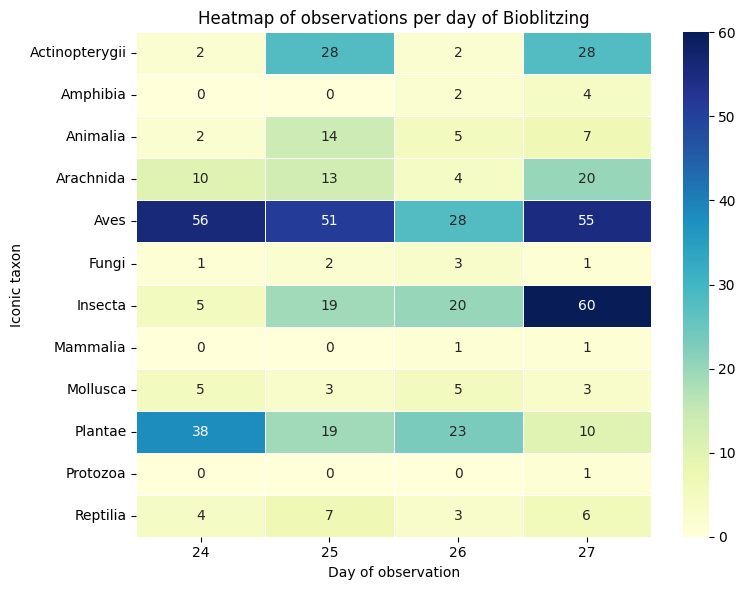
Top observers
top_obs = project_obs.groupby(['user_id','user_name']).agg({
'uuid': 'count',
'species_guess': ['count',pd.Series.nunique]})top_obs.loc[top_obs['uuid']['count']>10]| uuid | species_guess | |||
|---|---|---|---|---|
| count | count | nunique | ||
| user_id | user_name | |||
| 74355 | JR Ferrer-Paris | 93 | 83 | 62 |
| 1147408 | Andy Tuckey | 22 | 22 | 22 |
| 1453488 | Shannon Tushingham | 26 | 26 | 22 |
| 1771883 | Graham Winterflood | 11 | 11 | 9 |
| 4957699 | Sacha | 33 | 33 | 33 |
| 5414239 | SkippingShoe | 12 | 12 | 12 |
| 6287892 | Dr Spoodle Doodle | 14 | 14 | 10 |
| 6364733 | jimchurches | 40 | 40 | 33 |
| 8277420 | Matt Boyle | 16 | 16 | 15 |
| 9125842 | JimsWildLife | 56 | 50 | 35 |